Research scientists and clinicians involved : Amélie Bonnefond, Dominique Eladari, and Philippe Froguel
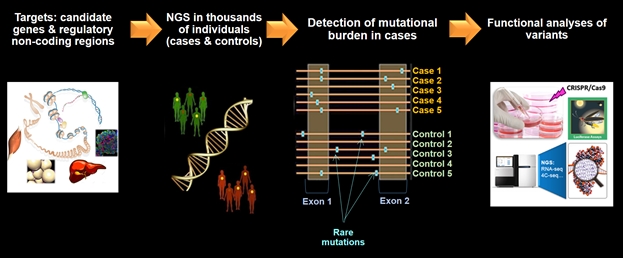
In common forms of type 2 diabetes and associated disorders, genome-wide association studies (GWAS) based on DNA microarrays used in large-scale population studies and cohorts, have demonstrated their polygenic nature, and have enabled the identification of hundreds of single nucleotide polymorphisms (SNPs) associated with disease risk. Despite this huge success, there has been a considerable gap between the knowledge of the genetic contribution of these SNPs and the understanding of how these genetic variants physiologically impact the disease: indeed, association does not mean causality. These GWAS-identified SNPs are frequent (with a minor allele frequency [MAF] >1%), mostly lie in intergenic or intronic regions and their individual effect on the disease risk is typically modest. All common SNPs identified by GWAS have explained less than 20% of disease heritability so far. The functional translation of these signals has been frustrating as the current functional assays (mostly based on cell or animal models) still lack sensitivity required for such subtle effects. For several years now, our team has investigated the contribution of rare DNA variants to the risk of complex forms of type 2 diabetes and associated disorders; this research has enabled insights into pathophysiology, along with putative identification of novel drug targets.
Related major publications of the team :
– Type 2 diabetes-associated variants of the MT2 melatonin receptor affect distinct modes of signaling. Karamitri A, Plouffe B, Bonnefond A, Chen M, Gallion J, Guillaume JL, Hegron A, Boissel M, Canouil M, Langenberg C, Wareham NJ, Le Gouill C, Lukasheva V, Lichtarge O, Froguel P, Bouvier M, Jockers R. Sci Signal. 2018 Aug 28;11(545).
– The Difficult Journey from Genome-wide Association Studies to Pathophysiology: The Melatonin Receptor 1B (MT2) Paradigm. Bonnefond A, Karamitri A, Jockers R, Froguel P. Cell Metab. 2016 Sep 13;24(3):345-347.
– Contribution of the low-frequency, loss-of-function p.R270H mutation in FFAR4 (GPR120) to increased fasting plasma glucose levels. Bonnefond A, Lamri A, Leloire A, Vaillant E, Roussel R, Lévy-Marchal C, Weill J, Galan P, Hercberg S, Ragot S, Hadjadj S, Charpentier G, Balkau B, Marre M, Fumeron F, Froguel P. J Med Genet. 2015 Sep;52(9):595-8.
– Interaction between GPR120 p.R270H loss-of-function variant and dietary fat intake on incident type 2 diabetes risk in the D.E.S.I.R. study. Lamri A, Bonnefond A, Meyre D, Balkau B, Roussel R, Marre M, Froguel P, Fumeron F; D.E.S.I.R. Study Group. Nutr Metab Cardiovasc Dis. 2016 Oct;26(10):931-6.
– Dysfunction of lipid sensor GPR120 leads to obesity in both mouse and human. Ichimura A, Hirasawa A, Poulain-Godefroy O, Bonnefond A, Hara T, Yengo L, Kimura I, Leloire A, Liu N, Iida K, Choquet H, Besnard P, Lecoeur C, Vivequin S, Ayukawa K, Takeuchi M, Ozawa K, Tauber M, Maffeis C, Morandi A, Buzzetti R, Elliott P, Pouta A, Jarvelin MR, Körner A, Kiess W, Pigeyre M, Caiazzo R, Van Hul W, Van Gaal L, Horber F, Balkau B, Lévy-Marchal C, Rouskas K, Kouvatsi A, Hebebrand J, Hinney A, Scherag A, Pattou F, Meyre D, Koshimizu TA, Wolowczuk I, Tsujimoto G, Froguel P. Nature. 2012 Feb 19;483(7389):350-4.
– Rare MTNR1B variants impairing melatonin receptor 1B function contribute to type 2 diabetes. Bonnefond A, Clément N, Fawcett K, Yengo L, Vaillant E, Guillaume JL, Dechaume A, Payne F, Roussel R, Czernichow S, Hercberg S, Hadjadj S, Balkau B, Marre M, Lantieri O, Langenberg C, Bouatia-Naji N; Meta-Analysis of Glucose and Insulin-Related Traits Consortium (MAGIC), Charpentier G, Vaxillaire M, Rocheleau G, Wareham NJ, Sladek R, McCarthy MI, Dina C, Barroso I, Jockers R, Froguel P. Nat Genet. 2012 Jan 29;44(3):297-301.