Research scientists and clinicians involved : Christophe Breton
a/ Identification of epigenomic targets during development.
Worldwide lifestyle changes, particularly excessive consumption of energy-dense foods and insufficient physical activity are driving forces of the global epidemic of obesity and metabolic complications. The Developmental Origin of Health and Disease (DOHaD) hypothesis suggests that adverse environments during either in utero or in the early postnatal development periods, result in long-term deleterious programming of metabolism.
Christophe Breton (PU in Physiology at Lille University) joined our team in 2020. His team previously reported in rodents that, maternal high-fat diet during gestation and lactation programs morphological and transcriptional alterations of adipose tissue in adult obesity-prone offspring via epigenetic modifications during development, in particular methylation and hydroxymethylation of specific/targeted sequences of the DNA.
Through the use of rodent models, they have deciphered that specific epigenetic marks are programmed by maternal obesity, which is associated to impaired glucose tolerance, insulin resistance and deterioration of insulin secretion with altered specific gene expression patterns in future adult T2D-prone offspring. In rodents, the maturation of adipose tissue is completed during gestation and lactation.
Thus, we postulate that adipose cells may represent prime targets of metabolic programming, and that maternal obesity may cause insulin resistance in offspring via epigenetically linked molecular mechanisms occurring early in life.
Using a mouse model of maternal high-fat diet-induced obesity, our aims are :
- 1) to analyze the architecture (i.e. size, vascularisation, proliferation, apoptosis, remodeling) and the function (i.e. adipokine synthesis and secretion, insulin signaling, lipogenesis and lipolysis) of adipocytes in offspring at different postnatal stages from lactation to adulthood;
- 2) to simultaneously analyze transcriptome profiles using RNA-seq, followed by molecular pathway bioinformatics analysis to determine the pathways targeted by the programming of adipose tissue;
- 3) to correlate the transcriptional profile changes of gene expression with DNA (hydroxy)methylation and histone modifications (i.e. active marks H3K27ac/H3K4me1 and inactive mark H3K27me3 that are mainly found in enhancer/promoter regions) in a epigenome-wide manner using (h)MeDIP-seq and ChIP seq approaches, respectively. In addition to the discovery of novel targets of programming, this project may constitute the first step towards developing strategies of intervention (i.e. epigenetic manipulations with small molecules) during the perinatal period to alleviate deleterious programming effects of maternal obesity on offspring’s adipose tissue via epigenome changes.
b/ Fighting insulin resistance through the browning of adipose tissue.
In rodents, genes that prevent fat accumulation and protect against diet-induced obesity (DIO) and T2D, such as E2f1 and Sertad2 (TRIP-Br2), improve insulin action in most metabolically active organs including adipose tissue.
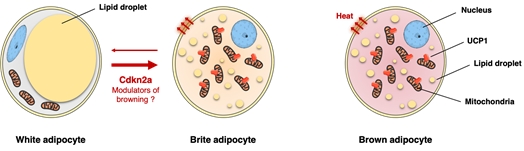
Adipose tissues include white (WAT), brown (BAT) and beige adipocytes. White adipocytes store excess energy as fat and release free fatty acids (FFAs) as energy substrate. The catabolic capability of BAT and beige adipocytes to burn fat contributes to reduce circulating FFAs. In obesity, elevated circulating FFAs are associated to insulin resistance and T2D. Since the energy balance is modulated by non-shivering thermogenesis, increasing energy expenditure by BAT activation or by promoting browning of WAT have recently emerged as new putative therapy for T2D.
In humans there are also arguments for a physiological role of brown/beige adipocytes to enhance glucose tolerance and insulin sensitivity. Therefore, the elucidation of the molecular pathways involved in beige adipocyte biogenesis would represent a first step towards innovative treatments (Figure 3).
Figure 3: the browning process as a new strategy to fight against insulin resistance and obesity.
As mentioned above, we demonstrated that the genetic deletion of Cdkn2a promotes energy expenditure and improves insulin sensitivity during diet-induced obesity through the control of beige adipocyte development. These data suggested that Cdkn2a plays a key role in energy metabolism by regulating a white-to-brown fat transition.
In addition, we observed that CDKN2A levels were increased in human obese adipocytes, further reinforcing CDKN2A as a potential target in obesity and T2D.
In collaboration with the team of C Dani, we are currently deciphering the molecular signature following the reprogramming of brown-like adipocytes derived from CDKN2A–deficient human Induced pluripotent stem cells. Using human induced pluripotent stem cells (hIPSCs) differentiated into brown-like adipocytes, we will identify the signaling pathways that contribute to this metabolic effect using genomic (RNA-seq/ChIP-seq) and whole kinome approaches (Pamgene.com) developed by our team.
Our second aim is to evaluate the metabolic effects of brown-like reprogrammed hIPSCs transplanted in mouse models of T2D and obesity. Our current results show that UCP1 levels are increased in siCDKN2A-hIPSCs differentiated brown-like cells compared to control cells. This suggests that these cells might be relevant to correct metabolic parameters during obesity.
Related major publications :
– Lecoutre S., Oger F., Pourpe C., Butruille L., Marousez L., Dickes-Coopman A., Laborie C., Guinez C., Lesage J., Vieau D., Junien C., Eberlé D., Gabory A., Eeckhoute J., Breton C. (2017). Maternal obesity programs increased leptin gene expression in rat male offspring via epigenetic modifications in a depot-specific manner. Mol. Metab. 6(8): 922-930.
– Lecoutre S., Pourpe C., Butruille L., Marousez L., Laborie C., Guinez C., Lesage J., Vieau D., Eeckhoute J., Gabory A., Oger F., Eberlé D., Breton C. (2018). Reduced PPAR expression in adipose tissue of male rat offspring from obese dams is associated with epigenetic modifications. FASEB 32(5): 2768-2778.
– Lecoutre S., Petrus P., Rydén M., Breton C. (2018). Transgenerational epigenetic mechanisms in adipose tissue development. Trends Endocrinol. Metab. 29: 675-685.
– Rabhi N., Hannou S.A., Gromada X., Salas E., Yao X., Oger F., Carney C., Lopez-Mejia I.C., Durand E., Rabearivelo I., Bonnefond A., Caron E., Fajas L., Dani C., Froguel P., Annicotte J.S. (2018). Cdkn2a deficiency promotes adipose tissue browning. Mol. Metab. 8:65-76.
– Kahoul Y., Oger F., Montaigne J., Froguel P., Breton C*., Annicotte J.S.* (2020). Emerging roles for the INK4a/ARF (CDKN2A) locus in adipose tissue: Implications for obesity and type 2 diabetes. Biomolecules. 10(9): 1350 (*co-last author).