Genomic plateform
The LIGAN PM-genomics platform (Lille Integrated Genomics Network for Advanced Personnalized medicine) aims to provide public and private research teams a complete set of tools and skills to simplify the study of genetics, genomics and transcriptomics. Our integrated solutions are designed to solve the challenges of genetic analysis and accelerate the pace of discovery. This labeled IBISA platform includes high quality technologies able to serve a large rank of projects (medium / high speed) of genotyping, sequencing, bioinformatics and bio-statisticals analyzes.
Service
The experience of a team for the development and support for transcriptomics and genomics projects, the choice of the appropriate technologies for these projects, training and technology development based on the novelties of the market. The technical staff ensures the proper functioning of equipment and organizes our activity according to the research projects and the platform of services.
Equipments and services
Our laboratory is equipped with technologies dedicated to the most efficient and innovative genomics. We are thus able to answer to all requests in this area, from robotic preparation of libraries for the high-throughput sequencing, including exomes sequencing projects, to targeted, genome and also metagenomic sequencing projects, de novo sequencing or mutation detection. Our expertise extends to the bioinformatics analyses and / or statistical data. UMR1283 publications associate to the equipments.

Equipments dedicated to Sequencing and NGS
- Hiseq 2500, Hiseq 4000, Nextseq 500, Miseq, CBot – Illumina
|
Illumina offers innovative next-generation sequencing (NGS) platforms that deliver industry-leading data quality and accuracy, at an unparalleled scale.NGS involves rapid sequencing of large DNA stretches spanning entire genomes. The latest Illumina DNA sequencers enable a wide variety of uses, and can produce gigabases to over a terabase of data in a single sequencing run.Common DNA Sequencing Applications:Illumina sequencing enables a broad range of applications, such as whole-genome and de novo sequencing, targeted region sequencing, mutation discovery, and identification of copy number variations. Common DNA sequencing applications include:Whole Genome Sequencing: Characterize entire genomes of any size and complexity.
- Exome sequencing: Sequence protein coding regions, as a cost-effective alternative to WGS.
- Targeted sequencing: Sequence specific genes or other regions of interest.
- De novo sequencing: Sequence and assemble novel genomes.
- Mitochondrial sequencing: Detect mitochondrial disease-associated mutations.
- Transcripome sequencing : transcriptome analysis
- Epigenetics: methyl sequencing
- Metagenomics: sequencing of 16S / 18S unti to characterize bacterial populations
Miseq : dedicated to small genome, amplicon and targeted gene panel sequencing
Nextseq500 and Hiseq 2500 and Hiseq 4000: dedicated to genome, exome, transcriptome sequencing |
    |

|
The Covaris E220 is a targeted ultrasound device designed for the treatment of samples of many biological and chemical applications, including DNA and shear chromatin tissue homogenization, cell lysis, dissolution compound, and micronization particles.
The use of acoustic energy (Adaptive Focused Acoustics ™) allows processing of 1-96 samples with precision and accuracy to samples of variable volumes.
|
 |

- Star – Hamilton & Bravo – Agilent
|
The Star (Hamilton) workstation and the Bravo Automated Liquid Handling Platform (Agilent) are liquid Handling workstations providing reproductibility and flexibility. Both dedicated to NGS libraries preparation, these equipments perform a large panel of automation protocols for Illumina Sequencing as libraries TruSeq DNA and RNA, Kapa DNA, Haloplex (Agilent), and WES or targeted captures (Nimblegen Roche, Sureselect Agilent) …
|

 |

- Pyromark Q24 advanced – QIAGEN
 |
PyroMark Q24 Advanced has improved Pyrosequencing® technology and provides even better real-time, sequence-based detection and quantification than before. PyroMark Q24 Advanced features advanced technology, software, and chemistry and is highly suited for analyzing any kind of sequence variation, particularly DNA methylation at CpG or CpN sites. The new CpN mode of PyroMark Q24 Advanced now enables methylation analysis of cytosine residues that are not part of CpG sites.
PyroMark Q24 Advanced enables improved methylation quantification in long sequence runs at any sequence position. Previously, analysis of methylation sites further away from the sequencing primer could be uncertain, but now with longer read lengths and higher accuracy, methylation quantification is highly reliable throughout the entire sequencing run.
|

|
Through a combination of Fluidigm’s exclusive integrated fluidic circuits (IFCs), the Access Array creates amplicon libraries using a unique tagging protocol, in which primers attach sample-specific barcode sequences and sequencer-specific tags to each PCR product. Multiplex primers build libraries of 480 amplicons per sample, and barcode sequences let you run up to 384 samples in one multiplex sequencing run.
|
 |

- SPRIworks SPRI-TE – Beckman
|
The SPRI-TE is an automated sample preparation for next generation sequencing Illumina or Roche. Based on magnetic bead purification technology (SPRI), it allows a differential sizing fragment and process up to 10 libraries in 5 hours.
|
 |

 |
The RainDance ThunderStorm System is a fully automated, high-throughput NGS content enrichment solution that enables researchers to automatically process up to 96 samples in 48 hours and to access up to eight different primer panels. The ThunderStorm System is based on RainDance’s proven single-molecule picodroplet PCR technology that generates millions of unique picodroplet PCR reactions, enabling scientists to target up to 20,000 genomic loci in a single sample. The system is compatible with all NGS platforms (Illumina sequencers, Roche sequencers, Life Technologies sequencers, and PacBio RS).
|

- Bioanalyzer 2100 – Agilent
 |
The Agilent 2100 bioanalyzer is the industry standard for RNA sample QC and has replaced labor-intensive gel electrophoresis for this application. It is also rapidly replacing gel electrophoresis for DNA fragment analysis and SDS-PAGE analysis of protein samples.
|

 |
The GS Junior System brings the power of NGS-454 Sequencing Systems directly to the laboratory benchtop. This system uses GS Junior Titanium chemistry; one run delivers an output of 35 Mbases with 400 bp reads in ten hours. The GS junior allows us a wide range of sequencing applications: amplicon sequencing, sequence capture, whole genome sequencing, metagenomics and transcriptome sequencing.
|

Equipments dedicated to genotyping, ddPCR, QPCR, proteomic, and expression studies
|
The iScan array scanner enables a wide variety of applications for superior genetic analysis studies, including:
- Genotyping : Illumina genotyping microarrays allow for powerful genome-wide association studies. Genotyping applications include:
- SNP genotyping and CNV analysis
- Linkage analysis
- Targeted genotyping
- Gene Expression Analysis : Gene expression arrays provide a global view of gene activity within cells. BeadArray technology enables robust and accurate measurement with wide dynamic range for large gene expression studies.
- FFPE Sample Analysis : Formalin-fixed, paraffin-embedded (FFPE) samples are preserved tissue samples that generally contain partially degraded RNA, making transcription analysis more challenging. Illumina offers solutions to provide high-quality array data from degraded RNA samples.
- Methylation Analysis : Epigenetic arrays let you quantitatively scan for methylation sites across the genome, utilizing high-multiplex assays for maximum throughput. Our Infinium Methylation Assay covers CpG islands, miRNA promoter regions, and other areas important in epigenetic regulation.
|
 |

|
Odyssey is an infrared imaging system which enables precise quantification over a range of about 4 log, and an increase of the sensitivity due to the absence of background noise.
Odyssey is equipped with two lasers, allowing detection simultaneously on two wavelengths (around 700 and 800 nm).
Immediate applications are standardizing results on the same matrix with the housekeeping gene, or the simultaneous detection of a protein and its phosphorylated form.
Finally, in addition to the “classic” western, the Odyssey can be used for many other applications like “in-cell” immunodetection experiments to quantify a protein on fixed cells in 24 or 96 micro-wells. This allows the study of the signaling pathways and screening without artifact due to cell lysis.
We can also count directly cells with a nuclear marker like DRAQ5.
|
 |

 |
The GloMax®-Multi Detection System can function as a luminometer (luciferase activity measurements) or a photometer (protein concentration quantification) in a 96 well plate format. Fast measurements, easy to use. Files can be exported in a .csv format.
|

- QX200 Droplet digital PCR system – BIORAD
|
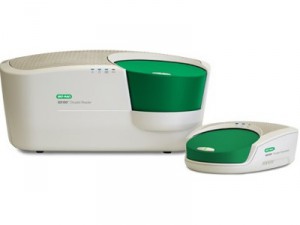
Droplet Digital PCR (ddPCR™) System provides absolute quantification of target DNA or RNA molecules for EvaGreen or probe-based digital PCR applications. The emerging applications are genomic alterations as gene copy number variation, NGS libraries quantification, absolute quantification, detection of rare sequences and rare mutations, gene expression and microRNA analysis, single cell analysis. The QX200 droplet generator partitions samples into 20,000 droplets. After PCR, droplets are read on a QX200 droplet reader in two hours, which counts the fluorescent positive and negative droplets to calculate target DNA concentration.
|
 |

 |
The integrated fluidic circuits (IFCs) allow automating PCR reactions in nanoliter volumes, so you use less sample and reagent. The Biomark HD system runs in either real-time or end-point read modes, bringing flexible, efficient and economical PCR solutions to a range of applications.In the lab we use 4 IFC :- 96.96 : 96 targets against 96 samples at once.- 48.48 : 48 targets against 48 samples at once.- 192.24 : 24 targets against 192 samples at once.– FLEXsix : six partitions (12 assays by 12-samples) that can each be run separately or together.
With these 4 IFC, we can do Gene expression and Genotyping studies.
We also study at the lab the polymorphism : Copy number variation (CNV).
The Digital Array IFC works by partitioning a single sample into 765 or 770 individual 6 nL real-time qPCR reactions.
The concentration of any sequence in a DNA sample (copies/μL) can be calculated using the number of positive chambers that contain at least one copy of that sequence.
For this type of study we have 2 types of IFC :
- – 765 : 12 samples into 765 chambers
- – 770 : 48 samples into 770 chambers
|

- nCounter analysis – Nanostring
 |
The nCounter® Analysis System from NanoString Technologies offers a simple, cost-effective way to profile hundreds of mRNAs, microRNAs, or DNA targets simultaneously with high sensitivity and precision. Digital detection of target molecules and high levels of multiplexing eliminate the compromise between data quality and data quantity, producing gold-standard sensitivity and reproducibility for studies of hundreds of targets. NanoString’s system uses molecular “barcodes” to detect and count hundreds of unique transcripts in a single reaction. Unlike other methods, the protocol does not include any amplification steps that might introduce bias to the results.
The systeme is divided in two parts:
- The nCounter Prep Station:. It processes nucleic acid samples post-hybridization to prepare them for data collection on the nCounter Digital Analyzer. Prior to placing samples on the Prep Station, samples are hybridized according to the nCounter protocol. On the deck of the Prep Station, hybridized samples are purified and subsequently immobilized in the sample cartridge for data collection. The Prep Station can process up to 48 samples in a 10-hour time span.
- The nCounter Digital Analyzer: it collects nucleic acid data by taking images of the immobilized fluorescent reporters in the sample cartridge with a CCD camera through a microscope objective lens. Images are processed internally and the data output files include the target identifier and count number along with a comprehensive tally of internal controls that allows each assay to be quantitative.
|

- Seahorse XFe24 – Seahorse Bioscience
|
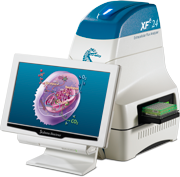
Seahorse XFe24 analyzers simultaneously measure mitochondrial respiration (oxygen consumption) and glycolysis (the release of H + ion) using non-invasive optical microsensors. Featuring a transitional microchamber the XFe24 allows for accurate measurements, non-invasive, non-destructive on 24 samples simultaneously with the sequential automatic injection of 4 drugs to specifically study different mitochondrial complexes, for example. One advantage is that the XFe24 is usable not only on monolayers of adherent primary cells (differentiated or not), cell lines, non-adherent cells, on isolated mitochondria but also on isolated pancreatic islets allowing accurate analysis of mitochondrial function in different tissue environments.
|
 |

- Viia7 Real Time PCR System – Life Technologies
|
The new ViiA™ 7 System is a 7th generation real-time PCR system that enables high-productivity qPCR through flexibility, and performance.
1°) Calculation of the absolute target quantity in samples with a standard curve. This method is able to determine the concentration of a target in samples and in a standard dilution series.
Spécific TaqMan Gene Expression Assays ou SYBR Green reagents can be used.
2°) Determination of the relative concentration of a sample or a target thanks to a standard curve and an endogenous control. The measures of concentrations are so normalized by using the endogenous control.
Relative Standard Curve experiments are commonly used with spécific TaqMan Gene Expression Assays to:
- Compare expression levels of a gene in different tissues.
- Compare expression levels of a gene in a treated sample and an untreated sample.
- Compare expression levels of wild-type alleles and mutated alleles.
- Analyze the gene expression changes over time under specific treatment conditions.
3°) Genotyping Experiments
Study of genetic variations (Single Nucleotid Polymorphism or Insertion/ Deletion) with spécific TaqMan Gene Expression Assays.
4°) Determination of the presence or the absence of the expression of a gene in a sample.
Performed PCR with two probes TaqMan specific to an internal control and to a gene of interest.
5°) Determination of the melting temperature of a nucleic acid sequence (TM). We performed with SYBR Green reagents a melting curve thanks to a temperature gradient at the end of the PCR. Determination of nonspecific PCR amplification.
Each study can be performed with 96 or 384 samples
|
 |

 |
TLDA means TaqMan Low Density Array and allows you to measure gene expression
using the comparative CT method of relative quantitation. It’s possible to study plenty of genes or miRNA on severals samples in a very small reaction volume (2 µl: 1 µl of sample and 1 µl of reaction mix).
Two types of 384 wells TaqMan Array cards are available:
– Customized cards containing the lyophilized Taqman assays of interest.
– Predefined cards for medical or scientific diagnostics.
|

Bioinformatics analysis and biostatistic analysis
The volume and quality of data produced by the new generation sequencers change regularly. Our laboratory regularly adapts and proposes the development of tools to answer and get the best for your data: assembly, transcriptome study, exomes and genomes analysis, statistics.
![]()
![]()
![]()
![]()
![]()
![]()
![]()
![]()
![]()
![]()
![]()
![]()
![]()
![]()
![]()
![]()
![]()
![]()
![]()

























